Converting the XFIT file into Crysfire format
Crysfire can import the following peak listing format.
EV = Bruker Socabim EVA format (EVA2Crys: Arie van der Lee) UD = Philips UDI format (UDI2Crys: Robin Shirley) WF = WinFit format (WF2Crys: Robin Shirley) XF = XFit format (XF2Crys: Robin Shirley)
The following assumes you have already created a Crysfire CDT file. Available, example tutorials for importing data into Crysfire are:
- Inputting a list of peaks / observations into Crysfire
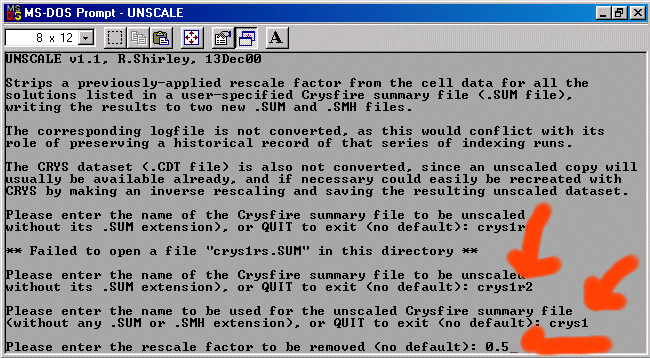
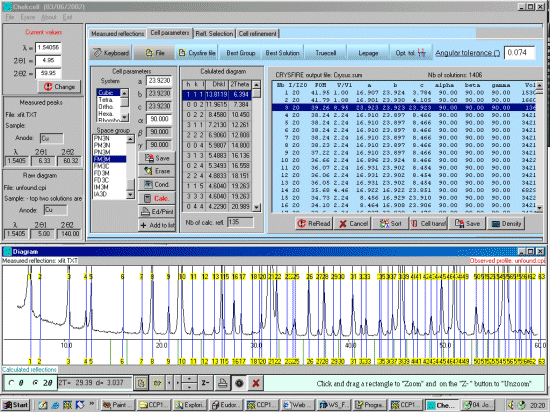
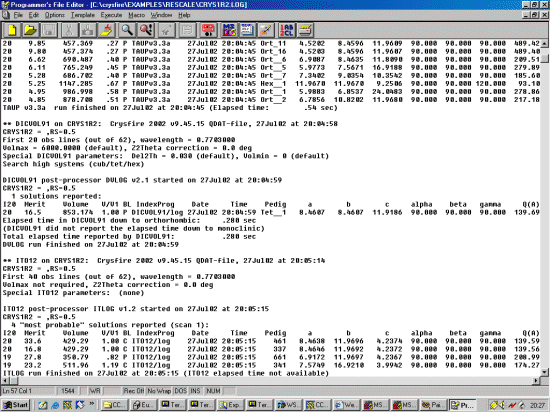
- Importing an XFIT Peak Listing into Crysfire and creating a CDT file (Winfit, Philips and Bruker files can also be imported into Crysfire)